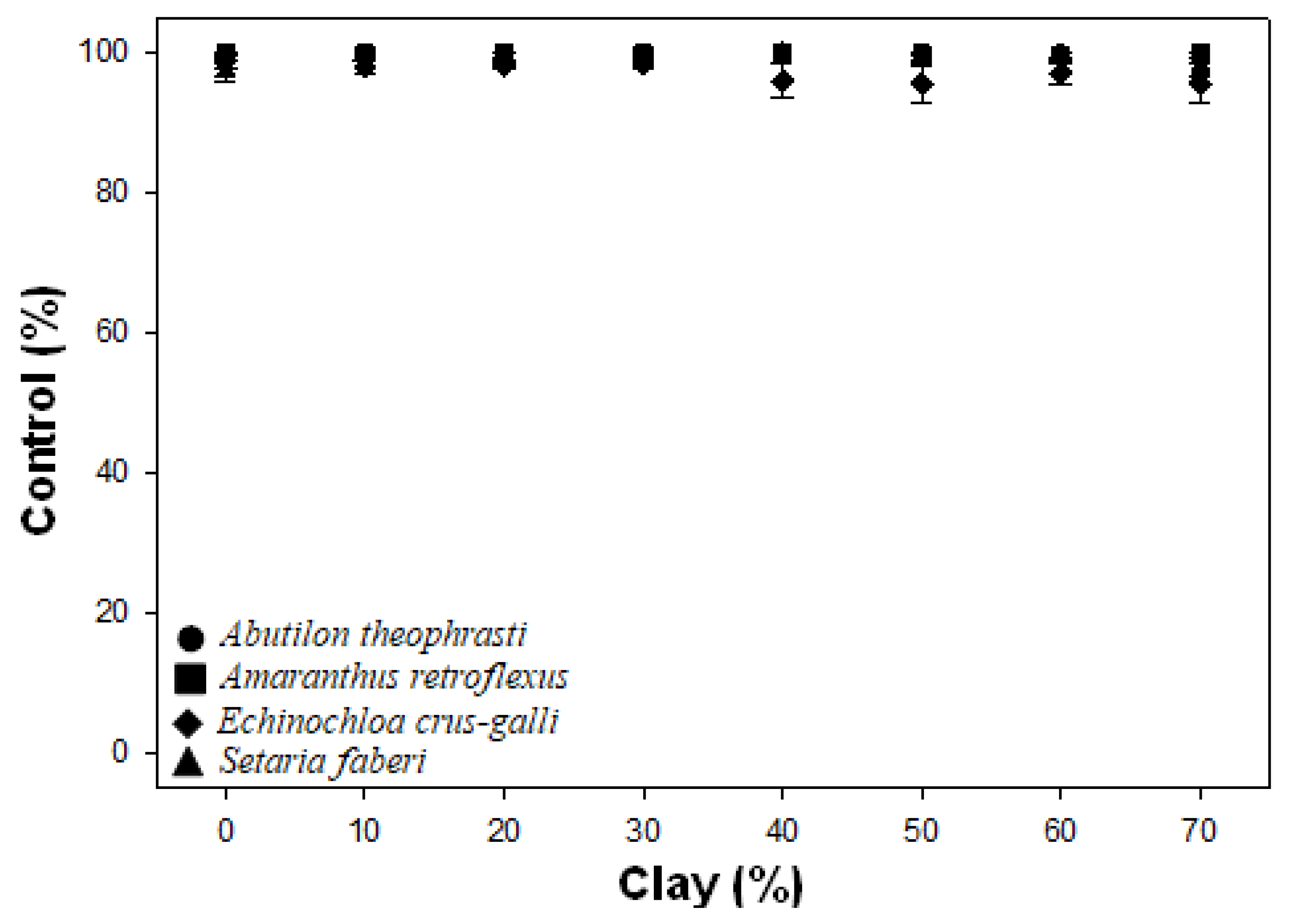
In the figure, the counts of the homopolymers of lengths 1-11 are plotted, with the Y-axis on a log scale. (The results for DH10B were identical to MG1655 and are therefore not shown), see the graph. I calculated the frequencies of homopolymers of all lengths present in both the Ion long reads dataset, the B13_328 dataset (100 bases run), the 454 dataset, and the E coli K12 MG1655 genome (both forward and reverse strands).

Due to the way the bases are read (basically not reading single bases, but the length of homopolymers). On the Ion Community forums, this peaks was discussed, but I couldn’t find a satisfactory explanation for it.įor problems with the data generated by both 454 and Ion Torrent, the usual suspect is homopolymers, consecutive runs of the same bases. But, they also show a strange peak around 400 bases. The Ion reads peak slightly lower than the 454 reads (241 and 253, respectively). I then took a closer look at the read length distribution of both datasets In a run I discussed earlier, I noticed a 32 base repeated flow order which presumably yields better results, but and this flow order was apparently not used for this run as well [thanks to Nils Homer for pointing out my oversight here, I’ve ordered new contact lenses…). The flow order used was reported as TACG all the way. Using my ‘2.5 bases per four flows’ calculation I described before, this translates in a potential 325 bases.
#Torrent sigmaplot 11 torrent
I randomly selected at the exact number of reads as are in the Ion Torrent B14_387 dataset.įirst, I looked into the sff file and noticed that the run had 520 flows, twice that was used earlier.
#Torrent sigmaplot 11 software
I used sff files that came with the software installation discs of newbler version 1.1, these represent a resequencing run on E coli K12 strain MG1655. For comparison, I generated a similar 454 GS FLX dataset, 350 000 reads with a peak length at 250 bases.

So I wondered what could be the problem with these reads, that presumably show good mapping accuracy, but perform not very well for assembly. On twitter, he wrote that the assembly programs MIRA and CLC Bio performed really bad with these reads. He reported, compared to the shorter Ion reads, much worse results for the longer reads. Nick Loman had a stab at using these reads for assembly with newbler, the program developed by 454 Life Sciences, on his blog.
And 454 reads were very useful at the time for de novo assembly (in fact, the only reads available for this purpose, obviously besides Sanger reads).
#Torrent sigmaplot 11 full
These accuracy measurements are logically based on alignment to a reference genome.īut what about de novo assembly? Thing is, the dataset presented, with a peak length of 241 (see below) and 350 000 read, is quite similar to what a full plate of GS FLX gave you in 2007 (peak length 250 bases, 400 000 reads).

There is an accompanying application note that brags about the read’s accuracy, especially over reads from the MiSeq platform. Life recently released ‘long’ IonTorrent reads (B14_387, resequencing of E coli strain DH10B, available through the Ion Community here). Trimming the ends off, however, only marginally improved de novo assembly of the reads using newbler. An analysis of the homopolymer distribution of the recently released ‘longer’ Ion Torrent reads indicates a possible significant over-calling of homopolymer lengths towards the ends of the reads.
